Research Directions
TorchXRayVision: A library for chest X-ray datasets and models. Including pre-trained models.
October 2021
TorchXRayVision is an open source software library for working with chest X-ray datasets and deep learning models. It provides a common interface and common pre-processing chain for a wide set of publicly available chest X-ray datasets. In addition, a number of classification and representation learning models with different architectures, trained on different data combinations, are available through the library to serve as baselines or feature extractors.
Project URL
Paper
GitHub
COVID-19 Image data collection
July 2020
Project Summary: To build a public open dataset of chest X-ray and CT images of patients which are positive or suspected of COVID-19 or other viral and bacterial pneumonias (MERS, SARS, and ARDS.). Data will be collected from public sources as well as through indirect collection from hospitals and physicians. All images and data will be released publicly in this GitHub repo.
Project URL
GitHub
Chester the AI Radiology Assistant
January 2019
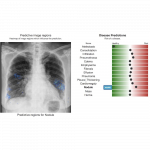
This is a web based (but locally run) prototype system for diagnosing chest X-ray images. The patient data remains on your computer and all computation occurs in your browser.
Project URL
Paper
GitHub
Gene Graph Convolutions
June 2018
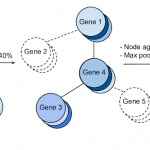
We discuss how gene-gene interaction graphs (same pathway, protein-protein, co-expression, or research paper text association) can be used to impose a bias on a deep neural network model similar to the spatial bias imposed by convolutions on an image. We find this approach provides an advantage for particular tasks in a low data regime but is very dependent on the quality of the graph used.
Paper
GitHub
Slides
Feature Hallucination in Medical Image Translation
May 2018
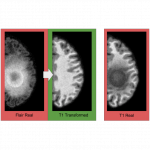
We show that cross domain image-to-image translation can be subject to bias due to matching the data distribution of the target domain. We specifically show results when using conditional GAN and CycleGAN.
Project URL
Paper
GitHub
Slides
Count-ception: Heterogeneous cell counting
March 2017
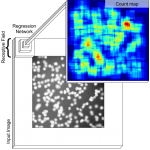
This work develops a method to count heterogeneous objects, such as
cell nuclei or sealions. We develop a deep learning based system to
that takes as input an image and returns a count of the objects inside
and justification for the prediction in the form of weak localization.
Paper
GitHub
Slides
Publications
Gifsplanation via Latent Shift: A Simple Autoencoder Approach to Counterfactual Generation for Chest X-rays
Released May 2021
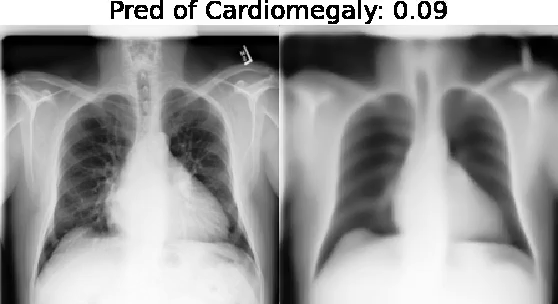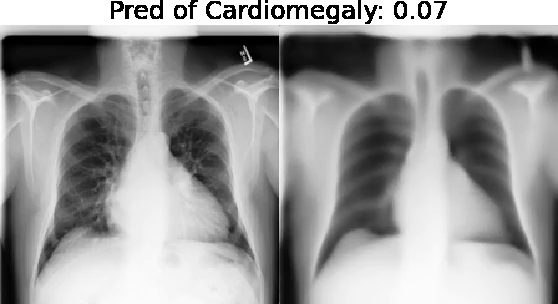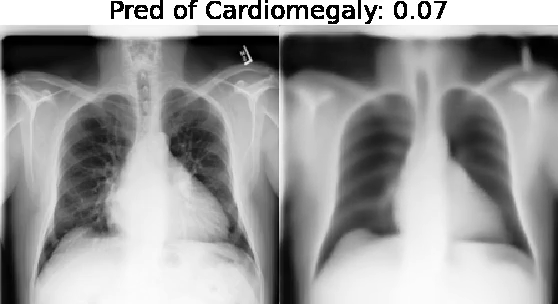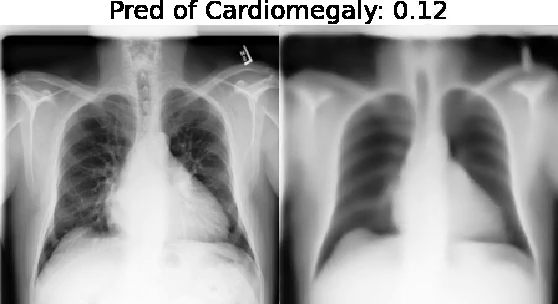 Motivation: Traditional image attribution methods struggle to satisfactorily explain predictions of neural networks. Prediction explanation is important, especially in medical imaging, for avoiding the unintended consequences of deploying AI systems when false positive predictions can impact patient care. Thus, there is a pressing need to develop improved models for model explainability and introspection.
Specific problem: A new approach is to transform input images to increase or decrease features which cause the prediction. However, current approaches are difficult to implement as they are monolithic or rely on GANs. These hurdles prevent wide adoption.
Our approach: Given an arbitrary classifier, we propose a simple autoencoder and gradient update (Latent Shift) that can transform the latent representation of a specific input image to exaggerate or curtail the features used for prediction. We use this method to study chest X-ray classifiers and evaluate their performance. We conduct a reader study with two radiologists assessing 240 chest X-ray predictions to identify which ones are false positives (half are) using traditional attribution maps or our proposed method.
Results: We found low overlap with ground truth pathology masks for models with reasonably high accuracy. However, the results from our reader study indicate that these models are generally looking at the correct features.
We also found that the Latent Shift explanation allows a user to have more confidence in true positive predictions compared to traditional approaches (0.15±0.95 in a 5 point scale with p=0.01) with only a small increase in false positive predictions (0.04±1.06 with p=0.57).
Motivation: Traditional image attribution methods struggle to satisfactorily explain predictions of neural networks. Prediction explanation is important, especially in medical imaging, for avoiding the unintended consequences of deploying AI systems when false positive predictions can impact patient care. Thus, there is a pressing need to develop improved models for model explainability and introspection.
Specific problem: A new approach is to transform input images to increase or decrease features which cause the prediction. However, current approaches are difficult to implement as they are monolithic or rely on GANs. These hurdles prevent wide adoption.
Our approach: Given an arbitrary classifier, we propose a simple autoencoder and gradient update (Latent Shift) that can transform the latent representation of a specific input image to exaggerate or curtail the features used for prediction. We use this method to study chest X-ray classifiers and evaluate their performance. We conduct a reader study with two radiologists assessing 240 chest X-ray predictions to identify which ones are false positives (half are) using traditional attribution maps or our proposed method.
Results: We found low overlap with ground truth pathology masks for models with reasonably high accuracy. However, the results from our reader study indicate that these models are generally looking at the correct features.
We also found that the Latent Shift explanation allows a user to have more confidence in true positive predictions compared to traditional approaches (0.15±0.95 in a 5 point scale with p=0.01) with only a small increase in false positive predictions (0.04±1.06 with p=0.57).
Paper
GitHub
Saliency is a Possible Red Herring When Diagnosing Poor Generalization
Released February 2021
Poor generalization is one symptom of models that learn to predict target variables using spuriously-correlated image features present only in the training distribution instead of the true image features that denote a class. It is often thought that this can be diagnosed visually using saliency maps. We study if this is correct. In some prediction tasks, such as for medical images, one may have some images with masks drawn by a human expert, indicating a region of the image containing relevant information to make the prediction. We study multiple methods that take advantage of such auxiliary labels, by training networks to ignore distracting features which may be found outside of the region of interest, on the training images for which such masks are available. This mask information is only used during training and has an impact on generalization accuracy depending on the severity of the shift between the training and test distributions. Surprisingly, while these methods improve generalization performance in the presence of a covariate shift, there is no strong correspondence between the correction of saliency towards the features a human expert have labelled as important and generalization performance. These results suggest that the root cause of poor generalization may not always be spatially defined, and raise questions about the utility of masks as "attribution priors" as well as saliency for explainable predictions.
Paper
GitHub
Predicting COVID-19 Pneumonia Severity on Chest X-ray with Deep Learning
Released May 2020
Purpose: The need to streamline patient management for COVID-19 has become more pressing than ever. Chest X-rays provide a non-invasive (potentially bedside) tool to monitor the progression of the disease. In this study, we present a severity score prediction model for COVID-19 pneumonia for frontal chest X-ray images. Such a tool can gauge severity of COVID-19 lung infections (and pneumonia in general) that can be used for escalation or de-escalation of care as well as monitoring treatment efficacy, especially in the ICU. Methods: Images from a public COVID-19 database were scored retrospectively by three blinded experts in terms of the extent of lung involvement as well as the degree of opacity. A neural network model that was pre-trained on large (non-COVID-19) chest X-ray datasets is used to construct features for COVID-19 images which are predictive for our task.
Results: This study finds that training a regression model on a subset of the outputs from an this pre-trained chest X-ray model predicts our geographic extent score (range 0-8) with 1.14 mean absolute error (MAE) and our lung opacity score (range 0-6) with 0.78 MAE. Conclusions: These results indicate that our model's ability to gauge severity of COVID-19 lung infections could be used for escalation or de-escalation of care as well as monitoring treatment efficacy, especially in the intensive care unit (ICU). A proper clinical trial is needed to evaluate efficacy.
Paper
GitHub
COVID-19 Image Data Collection
Released March 2020
Across the world's coronavirus disease 2019 (COVID-19) hot spots, the need to streamline patient diagnosis and management has become more pressing than ever. As one of the main imaging tools, chest X-rays (CXRs) are common, fast, non-invasive, relatively cheap, and potentially bedside to monitor the progression of the disease. This paper describes the first public COVID-19 image data collection as well as a preliminary exploration of possible use cases for the data. This dataset currently contains hundreds of frontal view X-rays and is the largest public resource for COVID-19 image and prognostic data, making it a necessary resource to develop and evaluate tools to aid in the treatment of COVID-19. It was manually aggregated from publication figures as well as various web based repositories into a machine learning (ML) friendly format with accompanying dataloader code. We collected frontal and lateral view imagery and metadata such as the time since first symptoms, intensive care unit (ICU) status, survival status, intubation status, or hospital location. We present multiple possible use cases for the data such as predicting the need for the ICU, predicting patient survival, and understanding a patient's trajectory during treatment.
Paper
GitHub
Quantifying the Value of Lateral Views in Deep Learning for Chest X-rays
Released February 2020
Most deep learning models in chest X-ray prediction utilize the posteroanterior (PA) view due to the lack of other views available. PadChest is a large-scale chest X-ray dataset that has almost 200 labels and multiple views available. In this work, we use PadChest to explore multiple approaches to merging the PA and lateral views for predicting the radiological labels associated with the X-ray image. We find that different methods of merging the model utilize the lateral view differently. We also find that including the lateral view increases performance for 32 labels in the dataset, while being neutral for the others. The increase in overall performance is comparable to the one obtained by using only the PA view with twice the amount of patients in the training set.
Paper
GitHub
Slides
On the limits of cross-domain generalization in automated X-ray prediction
Released February 2020
This large scale study focuses on quantifying what X-rays diagnostic prediction tasks generalize well across multiple different datasets. We present evidence that the issue of generalization is not due to a shift in the images but instead a shift in the labels. We study the cross-domain performance, agreement between models, and model representations. We find interesting discrepancies between performance and agreement where models which both achieve good performance disagree in their predictions as well as models which agree yet achieve poor performance. We also test for concept similarity by regularizing a network to group tasks across multiple datasets together and observe variation across the tasks.
Paper
GitHub
Chester: A Web Delivered Locally Computed Chest X-Ray Disease Prediction System
Released January 2019
In order to bridge the gap between Deep Learning researchers and medical professionals we develop a very accessible free prototype system which can be used by medical professionals to understand the reality of Deep Learning tools for chest X-ray diagnostics. The system is designed to be a second opinion where a user can process an image to confirm or aid in their diagnosis. Code and network weights are delivered via a URL to a web browser (including cell phones) but the patient data remains on the users machine and all processing occurs locally. This paper discusses the three main components in detail: out-of-distribution detection, disease prediction, and prediction explanation.
Paper
GitHub